Microarray and bioinformatics analysis of gene expression in experimental membranous nephropathy
PV. Hauser(1)*†, P. Perco(3,4,5)*, I. Mühlberger(5), J. Pippin(1), M. Blonski(1), B. Mayer(5), CE. Alpers(2), R. Oberbauer(3,4) SJ. Shankland(1)
1- Division of Nephrology & Hypertension, University of Washington, Seattle, WA, USA
2- Department of Pathology, University of Washington, Seattle, WA, USA
3- Department of Nephrology, KH Elisabethinen, Linz, Austria
4- Department of Nephrology, Medical University of Vienna, Austria
5- emergentec biodevelopment GmbH
†- currently at the Center for Molecular Biotechnology, University Torino, Torino, Italy
*- P. Hauser and P. Perco are co-first authors of this paper

Differentially expressed genes (DEGs)
List of 234 differentially regulated genes (>1.5 fold) in PHN induced rats on day three and or day six. Genes are annotated with NCBI Gene Symbols and grouped according to biological function. Starred (*) Gene Symbols mark predicted genes.
View PDF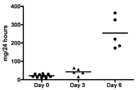
Proteinuria
Graph in figure 1 is showing total protein excretion of rats at the baseline (day 0) and after induction of disease (day 3 and day 6).
View PDF
List of over- and underrepresented ontology terms in the dataset
List of genes assigned to the over- or under-represented GO terms. Genes in the list are sorted according their cellular function. Significance levels are given as Bonferroni corrected p-values (p<0.005) following a chi-square test.
View PDF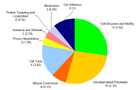
Enriched biological processes
Enriched biological processes and number of differentially expressed genes are given in figure 2. Numbers in brackets depict expected gene numbers according to the background distribution of all human genes.
View PDF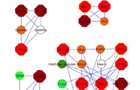
Protein interaction modules
Graphical visualization of the top four network modules. Node fill colour indicates the measured fold-change, where green represents down regulated genes (light green < -1.5, dark green < -2) and red represents upregulated genes (orange <1.5, red >1.5, dark red >2). The node’s border colour indicates the subcellular localization of the protein (yellow for membrane and blue for extra cellular). Hexagon shape of the node represents DEGs and circular shape represents direct interacting proteins.
View PDF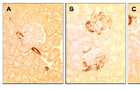
SM22 levels in PHN
(A) Staining for SM22 is absent in control glomeruli, but is detected in the blood vessels. (B) SM22 staining is detected in the glomerulus at day 3 of PHN, in a podocyte distribution. (C) SM22 staining is markedly increased at day 6 of PHN in podocytes. Staining was absent when the primary antibody was omitted (not shown).
View PDF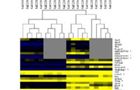
Heatmap showing expression levels of the 234 differentially expressed genes
Horizontal rows represent the expression levels of the 234 DEGs sorted by fold-change when comparing controls and treated animals. Arrays were clustered using average linkage distance and the Pearson correlation coefficient in the clustering algorithm. Gene Symbols marked with an asterisk depict predicted genes. Grey bars indicate non-significant expression changes.
View JPGOriginal data according to MiAME Guidelines
View PDFSample name:
RAE230A_d3_S1.CEL // rar-file
RAE230A_d3_S2.CEL // rar-file
RAE230A_d3_C3.CEL // rar-file
RAE230A_d3_C4.CEL // rar-file
RAE230A_d3_S5.CEL // rar-file
RAE230A_d3_S6.CEL // rar-file
RAE230A_d3_S7.CEL // rar-file
RAE230A_d3_C8.CEL // rar-file
RAE230A_d3_C9.CEL // rar-file
RAE230A_d3_C10.CEL // rar-file
RAE230A_d6_C1.CEL // rar-file
RAE230A_d6_C2.CEL // rar-file
RAE230A_d6_C3.CEL // rar-file
RAE230A_d6_C4.CEL // rar-file
RAE230A_d6_C5.CEL // rar-file
RAE230A_d6_S6.CEL // rar-file
RAE230A_d6_S7.CEL // rar-file
RAE230A_d6_S8.CEL // rar-file
RAE230A_d6_S9.CEL // rar-file
RAE230A_d6_S10.CEL // rar-file

